Key Word(s): Tuning, Hyperparameters, Random Forest
Title¶
Exercise 1: Bagging vs Random Forest (Tree correlation)
Description¶
How does Random Forest improve on Bagging?¶
The goal of this exercise is to investigate the correlation between randomly selected trees from Bagging and Random Forest.
Instructions:¶
- Read the dataset
diabetes.csvas a pandas dataframe, and take a quick look at the data. - Split the data into train and validation sets.
- Define a
BaggingClassifiermodel that usesDecisionTreClassifieras its base estimator. - Specify the number of bootstraps as 1000 and a maximum depth of 3.
- Fit the
BaggingClassifiermodel on the train data. - Use the helper code to predict using the mean model and individual estimators. The plot will look similar to the one given below.
- Predict on the test data using the first estimator and the mean model.
- Compute and display the validation accuracy
- Repeat the modeling and classification process above, this time using a
RandomForestClassifier.
Your final output should look something like this:¶
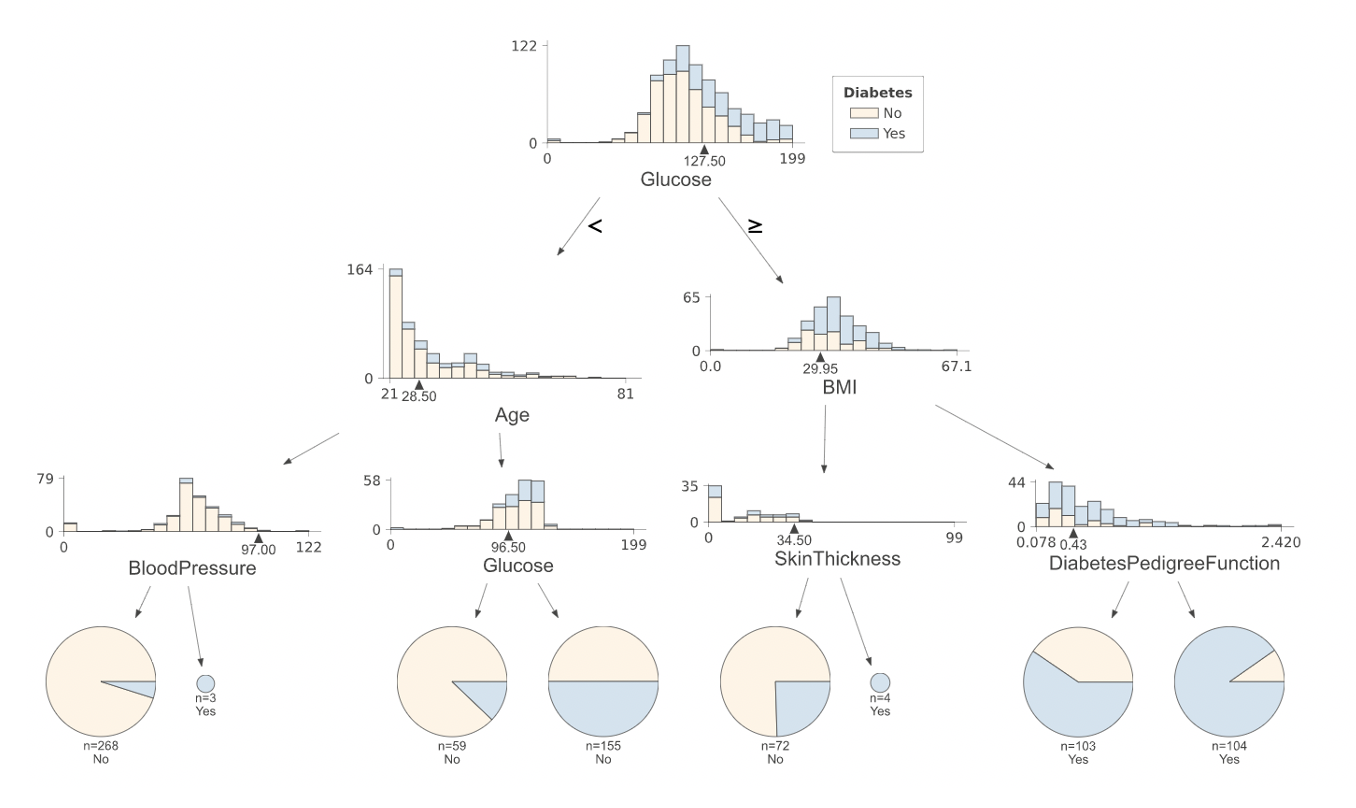
Hints:¶
sklearn.train_test_split() : Split arrays or matrices into random train and test subsets.
sklearn.ensemble.BaggingClassifier() : Returns a Bagging classifier instance.
sklearn.tree.DecisionTreeClassifier() : A Tree classifier can be used as the base model for the Bagging classifier.
sklearn.ensemble.andomForestClassifier() : Defines a Random forest classifier.
sklearn.metrics.accuracy_score(y_true, y_pred) : Accuracy classification score.
In [1]:
#!pip install -qq dtreeviz
import os, sys
sys.path.append(f"{os.getcwd()}/../")
In [2]:
# Import the main packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import BaggingClassifier
from sklearn.model_selection import train_test_split
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
from dtreeviz.trees import dtreeviz
%matplotlib inline
colors = [None, # 0 classes
None, # 1 class
['#FFF4E5','#D2E3EF'],# 2 classes
]
from IPython.display import Markdown, display
def printmd(string):
display(Markdown(string))
In [3]:
# Read the dataset and take a quick look
df = pd.read_csv("diabetes.csv")
df.head()
In [4]:
### edTest(test_assign) ###
# Assign the predictor and response variables.
# "Outcome" is the response and all the other columns are the predictors
X = __
y = __
In [5]:
# Fix a random_state and split the data
# into train and validation sets
random_state = 144
X_train, X_val, y_train,y_val = train_test_split(__,__,
train_size = 0.8,
random_state =random_state)
Bagging Implementation¶
In [6]:
# Define a Bagging classifier with randon_state as above
# and with a DecisionClassifier as a basemodel
# We fix the max_depth variable to 20 for all trees
max_depth = 20
# Set the maximum depth to be max_depth and use 100 estimators
n_estimators = 1000
basemodel = __(max_depth=__,
random_state=__)
bagging = BaggingClassifier(base_estimator=basemodel,
n_estimators=n_estimators)
# Fit the model on the training set
bagging.fit(__, __)
In [7]:
### edTest(test_bagging) ###
# We make predictions on the validation set
predictions = bagging.predict(X_val)
# compute the accuracy on the validation set
acc_bag = round(accuracy_score(predictions, y_val),2)
print(f'For Bagging, the accuracy on the validation set is {acc_bag}')
Random Forest implementation¶
In [8]:
# Define a Random Forest classifier with randon_state as above
# Set the maximum depth to be max_depth and use 100 estimators
random_forest = __(max_depth=max_depth,
random_state=random_state,
n_estimators=n_estimators)
# Fit the model on the training set
random_forest.fit(__, __)
In [9]:
### edTest(test_RF) ###
# We make predictions on the validation set
predictions = random_forest.predict(X_val)
# compute the accuracy on the validation set
acc_rf = round(accuracy_score(predictions, y_val),2)
print(f'For Random Forest, the accuracy on the validation set is {acc_rf}')
Visualizing the trees - Bagging¶
In [10]:
# Reducing the max_depth for visualization
max_depth = 3
basemodel = DecisionTreeClassifier(max_depth=max_depth,
random_state=random_state)
bagging = BaggingClassifier(base_estimator=basemodel,
n_estimators=1000)
# Fit the model on the training set
bagging.fit(X_train, y_train)
# Selecting two trees at random
bagvati1 = bagging.estimators_[0]
bagvati2 = bagging.estimators_[100]
In [11]:
vizA = dtreeviz(bagvati1, df.iloc[:,:8],df.Outcome,
feature_names = df.columns[:8],
target_name = 'Diabetes', class_names= ['No','Yes']
,orientation = 'TD',
colors={'classes':colors},
label_fontsize=14,
ticks_fontsize=10,
)
printmd(' Bagging Tree 1
')
vizA
In [12]:
vizB = dtreeviz(bagvati2, df.iloc[:,:8],df.Outcome,
feature_names = df.columns[:8],
target_name = 'Diabetes', class_names= ['No','Yes']
,orientation = 'TD',
colors={'classes':colors},
label_fontsize=14,
ticks_fontsize=10,
scale=1.1
)
printmd(' Bagging Tree 2
')
vizB
Visualizing the trees - Random Forest¶
In [13]:
# Reducing the max_depth for visualization
max_depth = 3
random_forest = RandomForestClassifier(max_depth=max_depth, random_state=random_state, n_estimators=1000,max_features = "sqrt")
# Fit the model on the training set
random_forest.fit(X_train, y_train)
# Selecting two trees at random
forestvati1 = random_forest.estimators_[0]
forestvati2 = random_forest.estimators_[__]
In [14]:
vizC = dtreeviz(forestvati1, df.iloc[:,:8],df.Outcome,
feature_names = df.columns[:8],
target_name = 'Diabetes', class_names= ['No','Yes']
,orientation = 'TD',
colors={'classes':colors},
label_fontsize=14,
ticks_fontsize=10,
scale=1.1
)
printmd(' Random Forest Tree 1
')
vizC
In [15]:
vizD = dtreeviz(forestvati2, df.iloc[:,:8],df.Outcome,
feature_names = df.columns[:8],
target_name = 'Diabetes', class_names= ['No','Yes']
,orientation = 'TD',
colors={'classes':colors},
label_fontsize=14,
ticks_fontsize=10,
scale=1.1
)
printmd(' Random Forest Tree 2
')
vizD
Mindchow 🍲¶
- Change the
max_depthof Bagging and Random Forest to see different trees. Which one gives different trees? - Change the
max_featuresinRandomForestClassifierto 8. How is it affecting the correlation between the trees?